Latest recommendations

| Id | Title * | Authors * | Abstract * | Picture * | Thematic fields * | Recommender▲ | Reviewers | Submission date | |
|---|---|---|---|---|---|---|---|---|---|
04 Mar 2024
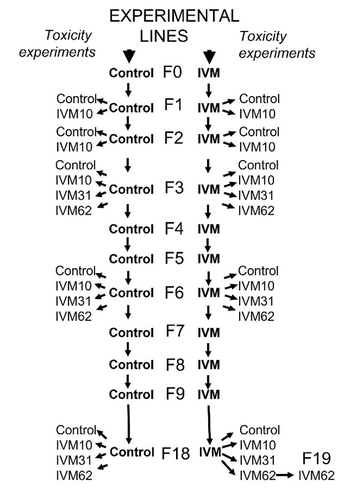
Ivermectin resistance in dung beetles exposed for multiple generationsDaniel Gonzalez Tokman, Antonio Arellano Torres, Fernanda Baena-Diaz, Carlos Bustos, Imelda Martinez M https://doi.org/10.1101/2023.05.08.539900Low potential of arthropod species to aquire resistance to invermectin drug could induce a risk of extinction in contaminated pasturesRecommended by Christian Mougin based on reviews by Marcel Amichot and 2 anonymous reviewers based on reviews by Marcel Amichot and 2 anonymous reviewers
For many decades, the macrocyclic lactone drug ivermectin is extensively used in veterinary medicine and agriculture, as well as human medicine. Residues of ivermectin excreted in cattle dung remain persistent in soils (Mougin et al., 2003), biologically active and threaten non-target soil and coprophagous organisms such as dung flies and beetles (Lumaret et al., 2012). Ivermectin affects highly beneficial and taxonomically diverse groups inhabiting dung pats, including flies, parasitic wasps, as well as coprophilus and predatory dung beetles (Villar et al., 2022). Ivermectin resistance is well document in insects, but it seems to take longer and to be less effective than resistance to insecticides or other antiparasitic drugs, because of different physiological mechanisms involved in resistance (Seaman et al., 2015). In that context, Gonzalez-Tokman et al. (2024) compared the reproductive success of a line of dung beetles (Euoniticellus intermedius, Scarabaeinae) exposed to a moderate concentration of invermectin during 18 generations, and a control line of beatles that was maintained free of antiparasitic drug. They carried-out toxicity experiments with increasing ivermectin concentrations to determine if sensitivity to ivermectin was reduced after some generations of exposure, possibly by acquiring resistance by means of transgenerational effects. Thus, dung beetles did not generate resistance to ivermectin after 18 generations of continuous exposure, and quantitative genetic analyses showed only low genetic variation in response to ivermectin. The results published by Gonzalez-Tokman et al. (2024) indicated a low potential of beetles for adaptation to the drug, and suggest for non-target invertebrate groups a possible risk of extinction in ivermectin-contaminated pastures. These effects can greatly impact grassland ecology, lower their quality and reduce the area available and palatable to livestock. References Mougin, C., Kollmann, A., Dubroca, J., Ducrot, P.-H., Alvinerie, M., Galtier, P., 2003. Fate of the veterinary medicine ivermectin in soil. Environ. Chem. Letters 1, 131-134. https://doi.org/10.1007/s10311-003-0032-9 Lumaret, JP., Errouissi, F., Floate, K., Römbke, J., Wardhaugh, K., 2012. A review on the toxicity and non-target effects of macrocyclic lactones in terrestrial and aquatic environments. Current Pharmaceutical Biotechnology 13(6), 1004-60. https://doi.org/10.2174/138920112800399257 Villar, D., & Schaeffer, D.J., 2022. Ivermectin use on pastured livestock in Colombia: parasite resistance and impacts on the dung community. Revista Colombiana De Ciencias Pecuarias, 36(1), 3-12. https://doi.org/10.17533/udea.rccp.v36n1a2 Seaman, J.A., Alout, H., Meyers, J.I., Stenglein, M.D., Dabiré, R.K., Lozano-Fuentes, S., Burton, T.A., 471 Kuklinski, W.S., Black, W.C., Foy, B.D., 2015. Age and prior blood feeding of Anopheles gambiae influences their susceptibility and gene expression patterns to ivermectin-containing blood meals. BMC Genomics 16, 797. https://doi.org/10.1186/s12864-015-2029-8 González-Tokman, D., Arellano-Torres, A., Baena-Díaz, F., Bustos, C., Martínez M., I., 2024. Ivermectin resistance in dung beetles exposed for multiple generations, bioRxiv ver. 3 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://doi.org/10.1101/2023.05.08.539900 | Ivermectin resistance in dung beetles exposed for multiple generations | Daniel Gonzalez Tokman, Antonio Arellano Torres, Fernanda Baena-Diaz, Carlos Bustos, Imelda Martinez M | <p>Ivermectin is an antiparasitic drug commonly used in cattle, that is excreted in dung, causing lethal and sub-lethal effects on coprophagous non-target fauna. Given that cattle parasites generate resistance to ivermectin, farmers have increased... |  | Ecosystem Health, Environmental pollution, Global changes, Terrestrial ecotoxicology | Christian Mougin | 2023-05-12 04:57:32 | View | |
22 Jul 2023
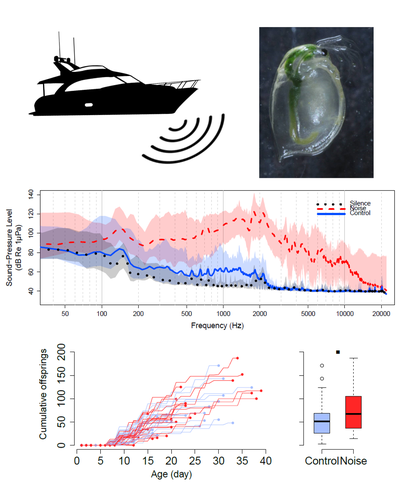
No evidence for an effect of chronic boat noise on the fitness of reared water fleasLoïc Prosnier, Emilie Rojas, Vincent Médoc https://doi.org/10.1101/2022.11.20.517267Noise impact in Daphnia magnaRecommended by Claudia Cosio based on reviews by Marie-Agnès Coutellec and 1 anonymous reviewerOur ability to anticipate and estimate how pollution affects biota is of paramount importance in the field of ecotoxicology. Impact of chemical pollution by metals, drugs or pesticides was widely studied in different species using acute and chronic scenarios. While environmental factors such as temperature are also often considered, noise is largely ignored in these models despite the knowledge of its detrimental effects in vertebrates. Studies of noise impacts included behavior and fitness endpoints and showed no effect to death depending on intensity, frequency and the distance from the noise source (Peng et al., 2015). Nonetheless, the impact of noise in biota is not well-understood, which impairs its effective mitigation. Noise or acoustic pollution due to boat traffic produce low-frequency stationary noise. It is a pervasive and ubiquitous pollutant found in aquatic ecosystems. In this context, Prosnier et al. (2023) addresses how intermittent and random noise impacted Daphnia magna, a representative of zooplankton model, widely used in ecotoxicology. Endpoints of lifespan and clonal offspring production were measured in the presence or absence of motorboat noises, in animals reared from birth to death. Noise consisted in a playlist of 15 sounds of motorboat recorded in the Grangent lake (Loire, France). Their intensity ranged from 0 to -25 dB Re 1 μPa by 5 dB to create 75 sounds from 103 to 150 dB RMS Re 1 μPa – a range of levels occurring in lakes. Treatment had no effect on analyzed endpoints, contrary to a continuous broadband noise (100-20,000 Hz) that caused higher survival and fecundity, and reduced speed of motion compared to control (Prosnier et al., 2022). Data point that temporal (continuous, regular, random) and frequency of noise are instrumental for its effects. References Peng, C., X. Zhao and G. Liu (2015). "Noise in the Sea and Its Impacts on Marine Organisms." Int J Environ Res Public Health 12(10): 12304-12323. https://doi.org/10.3390/ijerph121012304 Prosnier, L., E. Rojas and V. Médoc (2023). "No evidence for an effect of chronic boat noise on the fitness of reared water fleas." bioRxiv: 2022. ver. 4 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://doi.org/10.1101/2022.11.20.517267 Prosnier, L., E. Rojas, O. Valéro and V. Médoc (2022). "Chronic noise unexpectedly increases fitness of a freshwater zooplankton." bioRxiv: 2022. https://doi.org/10.1101/2022.11.19.517212 | No evidence for an effect of chronic boat noise on the fitness of reared water fleas | Loïc Prosnier, Emilie Rojas, Vincent Médoc | <p style="text-align: justify;">Among the numerous questions about human impacts on ecosystems, there is a growing interest for acoustic pollution. First studies on underwater acoustic pollution focused, and showed effects, on vertebrates’ behavio... |  | Aquatic ecotoxicology, Ecosystem Health, Environmental pollution, Global changes, Life History, Other | Claudia Cosio | 2022-12-08 17:23:07 | View | |
25 Sep 2023
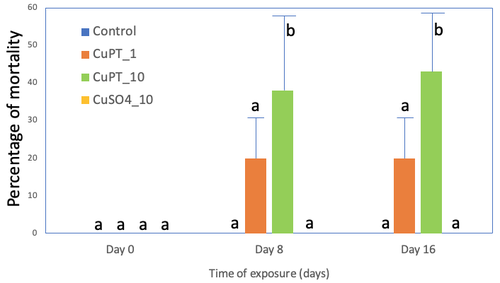
Characterization of the bioaccumulation and toxicity of copper pyrithione, an antifouling compound, on juveniles of rainbow troutCharlotte Bourdon, Jérôme Cachot, Patrice Gonzalez, Patrice Couture https://doi.org/10.1101/2023.01.31.526498Bioaccumulation and impact of copper pyrithione impact in juveniles of rainbow troutRecommended by Claudia Cosio based on reviews by Anne-Sophie Voisin and 1 anonymous reviewerOur ability to anticipate and estimate how pollution affects biota is intrumental in the field of ecotoxicology. Impact of chemical pollution by metals, drugs or pesticides was widely studied in different species using acute and chronic scenarios. Since the ban on tributyltin in antifouling paints, other copper (Cu)-based paints are on the market, including a new generation of booster biocides:metal pyrithiones such as Cu pyrithione (CuPT). Pyrithione acts as a Cu ionophore facilitating Cu transport across the membranes. Although some data show their occurrence in aquatic ecosystems and few studies on the toxicity of CuPT in fish are published, major gaps in knowledge remain about their toxicity and toxic pathway. Few studies were previously conducted in animals exposed to CuPT pointing to reprotoxicity, developmental malformation and mortality (Li et al. 2021, Mochida et al., 2011; Mohamat-Yusuff et al., 2018, Shin et al., 2022). However, its toxicokinetic and toxicodynamic remain to be characterized in details. In this context, Bourdon et al. (2023) compared in juveniles of rainbow trout (Oncorhynchus mykiss), the effects of exposure to CuPT and ionic Cu2+ at equivalent Cu2+ molar concentrations. Presented data allow to compare the toxicity threshold, the accumulation of Cu and mechanisms of toxicity of both compounds. Acute and chronic exposures showed a higher bioaccumulation of Cu in the gills, and a higher toxicity of CuPT than that of ionic Cu2+, e.g. mortality , transcription levels of genes related to oxidative stress, detoxification and Cu transport. Intriguingly, the activities of enzymatic biomarkers used as proxy of antioxidant capacity were not significantly altered, although Cu is generally expected to trigger oxidative stress. In conlusion, this study brings new knowledge pointing that the presence of CuPT in the environment could induce toxic effects in non-target species. Moreover, it support the need to study in detail the toxicity of Cu-based paints to adapt regulations concerning their use and release in aquatic environments. Because of its low solubility in water, CuPT is expected to adsorb to suspended matter and food pellets. Future research should study this route of exposure.
References Bourdon, C., Cachot, J., Gonzalez, P., Couture, P., 2023. Characterization of the bioaccumulation and toxicity of copper pyrithione, an antifouling compound, on juveniles of rainbow trout, bioRxiv ver. 3 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://doi.org/10.1101/2023.01.31.526498 Li, X., S. Ru, H. Tian, S. Zhang, Z. Lin, M. Gao and J. Wang, 2021. Combined exposure to environmentally relevant copper and 2,2′-dithiobis-pyridine induces significant reproductive toxicity in male guppy (Poecilia reticulata). Science of the Total Environment 797, https://doi.org/10.1016/j.scitotenv.2021.149131 Mochida, K., Amano, H., Onduka, T., Kakuno, A., Fujii, K., 2011. Toxicity and metabolism of copper pyrithione and its degradation product, 2,2’-dipyridyldisulfide in a marine polychaete. Chemosphere 82, 390–397, https://doi.org/10.1016/j.chemosphere.2010.09.074 Mohamat-Yusuff, F., Sarah-Nabila, Ab.G., Zulkifli, S.Z., Azmai, M.N.A., Ibrahim, W.N.W., Yusof, S., Ismail, A., 2018. Acute toxicity test of copper pyrithione on Javanese medaka and the behavioural stress symptoms. Marine Pollution Bulletin 127, 150–153, https://doi.org/10.1016/j.marpolbul.2017.11.046 Shin, D., Y. Choi, Z. Y. Soon, M. Kim, D. J. Kim and J. H. Jung, 2022. Comparative toxicity study of waterborne two booster biocides (CuPT and ZnPT) on embryonic flounder (Paralichthys olivaceus). Ecotoxicology and Environmental Safety 233, https://doi.org/10.1016/j.ecoenv.2022.113337 | Characterization of the bioaccumulation and toxicity of copper pyrithione, an antifouling compound, on juveniles of rainbow trout | Charlotte Bourdon, Jérôme Cachot, Patrice Gonzalez, Patrice Couture | <p>Since the global ban on tributyltin in antifouling paints in 2008 by the International Maritime Organization, new products have been developed and brought to the market. Among them, copper pyrithione (CuPT) is used, but its mechanisms of toxici... |  | Aquatic ecotoxicology, Bioassays, Biomarkers, Biomonitoring, Biotransformation, Environmental pollution | Claudia Cosio | Elise David, Anne-Sophie Voisin | 2023-02-01 15:23:44 | View |
22 Jul 2023

DRomics, a workflow to exploit dose-response omics data in ecotoxicologyMarie Laure Delignette-Muller, Aurélie Siberchicot, Floriane Larras, Elise Billoir https://doi.org/10.1101/2023.02.09.527852New features of DRomics workflow for improved analyze of dose-response omics data in ecotoxicologyRecommended by Claudia Cosio based on reviews by Jean Armengaud, Beatrice Gagnaire and Rebecca BeauvaisOur ability to anticipate and estimate how pollution affects components of ecosystems is of paramount importance in the field of ecotoxicology. Dose-response modeling is instrumental, as it allows deriving sensitivity thresholds used at the basis of regulatory risk assessment. In recent years, omics have highly influenced how the impacts of stressors are understood by revealing molecular changes at all levels of biota biological organization (Ebner et al., 2021). To allow analysis of omics data obtained using a typical dose-response design, DRomics a freely available tool for dose-response was proposed composed of both an R package and a free web application (Larras et al. 2018). Advances in this field depend both on theoretical concepts, technology and data integration. In this context, Delignette-Muller et al. (2023) address the question of how to better integrate omics information in dose-response questions. The paper lists previous possibilities of DRomics and presents new features. It is now able to handle all types of continuous omic and continuous non-omic data (e.g. growth data). This new version proposes new visualization tools, functional annotation and improved modeling workflow for a better robustness of analysis of data with few replicates. New features are meant to help for biological interpretation at the metabolic pathway level, to compare different measurements, biological materials or experimental settings. References Delignette-Muller, M. L., A. Siberchicot, F. Larras and E. Billoir (2023), DRomics, a workflow to exploit dose-response omics data in ecotoxicology. bioRxiv, 2023.2002.2009.527852, ver. 4 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://doi.org/10.1101/2023.02.09.527852 Ebner JN. (2021) Trends in the Application of "Omics" to Ecotoxicology and Stress Ecology. Genes, 12(10):1481. https://doi.org/10.3390/genes12101481 Larras F, Billoir E, Baillard V, Siberchicot A, Scholz S, Wubet T, Tarkka M, Schmitt-Jansen M and Delignette-Muller ML (2018). DRomics: a turnkey tool to support the use of the dose-response framework for omics data in ecological risk assessment. Environmental science & technology, 52(24):14461. https://doi.org/10.1021/acs.est.8b04752 | DRomics, a workflow to exploit dose-response omics data in ecotoxicology | Marie Laure Delignette-Muller, Aurélie Siberchicot, Floriane Larras, Elise Billoir | <p style="text-align: justify;">Omics technologies has opened new possibilities to assess environmental risks and to understand the mode(s) of action of pollutants. Coupled to dose-response experimental designs, they allow a non-targeted assessmen... |  | Aquatic ecotoxicology, Environmental risk assessment, Genetics / Genomics, Marine ecotoxicology, Microbial ecotoxicology, Modelling, Terrestrial ecotoxicology | Claudia Cosio | Rebecca Beauvais | 2023-02-17 15:39:03 | View |
02 May 2024

Maternal body condition affects the response of larval spined toads' faecal microbiome to a widespread contaminantSabrina Tartu, Nicolas Pollet, Isabelle Clavereau, Gauthier Bouchard, Francois Brischoux https://doi.org/10.1101/2023.12.18.572122Effects of AMPA on Bufo spinosus microbiotaRecommended by Marie-Agnès Coutellec based on reviews by Fabrice Martin-Laurent, Lauris Evariste and 1 anonymous reviewerThe overall pollution of air, water, and soil is currently recognized as one of the five main drivers of biodiversity loss (IPBES 2019). Among chemicals, pesticides play a significant role in this global crisis, as recently re-assessed at the scale of France (Pesce et al. 2023). In this context, although parent molecules are subject to national and international regulations, based on a priori ecological risk assessment (e.g., REACH) as well as monitoring in some environments (see e.g., pesticides classified in the priority list of substances by the Water Frame Directive), pesticide metabolites are rarely considered. In the case of the widely used herbicide glyphosate, a particular concern is rising about its primary metabolite, aminomethylphosphonic acid (AMPA), due to its persistence and overlooked toxicity. Amphibians are the most threatened class of vertebrates on earth, with two in every five species considered threatened with extinction (IUCN Red List). While this overall decline has multiple causes, the contribution of pesticides is suspected to be significant in some regions. In this context, Tartu et al. (2024) studied the effects of AMPA on the gut microbiota of the spined toad, Bufo spinosus. This work complements a previous study which showed embryo mortality, oxidative stress, deformities at hatching, and delayed development (Tartu et al. 2022). Using a common garden experiment based on populations from contrasted habitats (agricultural vs woodland, same as in the previous study), the authors captured breeding pairs and collected the eggs laid in the laboratory. These were exposed to 0.4 µg/L AMPA during embryonic and larval development. Individual microbiota was analysed non-invasively, i.e., using the faeces collected in treatment vessels. Bacterial biodiversity was genetically assessed (16S rRNA). The community biomass and taxonomic structure were analysed as a function of chemical treatment, mother and father body condition (fat vs thin), as well as population of origin. As a primary effect, AMPA reduced the microbial biomass. Furthermore, a significant interaction was detected between AMPA and mother condition on the community structure and composition. This alteration, observed in « fat » females only, was reflected through a significant decrease in Bacteroidota and a significant increase in Actinobacteriota (the latter being consistent with the ability of some species in this phylum to use AMPA as a source of phosphorus). The higher sensitivity of tadpoles from females in better condition seems counterintuitive, since better body condition is expected to be associated with higher fitness (and possibly higher ability to face chemical stress), the authors discuss this in the light of the literature (which shows that microbiome-fitness relationships are not often evidenced in natural populations), and hypothesize that these females in better conditions host a microbiota that may be more efficient, yet also more sensitive to AMPA. Not ruling out other possible factors ignored in their study, in particular genotypic effects, the authors further discuss the importance of maternally transmitted effects via the microbiota. Altogether, the results published by Tartu et al. (2024) provide important new findings on AMPA toxicity to amphibian microbiota, and also confirm the occurrence of vertical transmission of the microbiota from mother to progeny in this vertebrate class. References IPBES (2019). Global assessment report on biodiversity and ecosystem services of the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services. E. S. Brondizio, J. Settele, S. Díaz, and H. T. Ngo (editors). IPBES secretariat, Bonn, Germany. 1148 pages. https://doi.org/10.5281/zenodo.3831673 Pesce, S., Mamy, L., Sanchez, W., et al. (2023). Main conclusions and perspectives from the collective scientific assessment of the effects of plant protection products on biodiversity and ecosystem services along the land–sea continuum in France and French overseas territories. Environ Sci Pollut Res . https://doi.org/10.1007/s11356-023-26952-z Tartu, S., Renoirt, M., Cheron, M., Gisselmann, L.-L., Catoire, S., Brischoux, F. (2022). Did decades of glyphosate use have selected for resistant amphibians in agricultural habitats? Environ. Pollut. 310, 119823. https://doi.org/10.1016/j.envpol.2022.119823 Tartu, S., Pollet, N., Clavereau, I., Gauthier Bouchard, G., Brischoux, F. (2024). Maternal body condition affects the response of larval spined toads’ faecal microbiome to a widespread contaminant. bioRxiv, ver. 2 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://doi.org/10.1101/2023.12.18.572122
| Maternal body condition affects the response of larval spined toads' faecal microbiome to a widespread contaminant | Sabrina Tartu, Nicolas Pollet, Isabelle Clavereau, Gauthier Bouchard, Francois Brischoux | <p>Glyphosate’s primary metabolite, aminomethylphosphonic acid (AMPA), is the most detected pollutant in surface waters. Recent studies have raised concerns about its toxicity, yet underlying mechanisms remain poorly understood. A disruption of th... |  | Aquatic ecotoxicology, Environmental pollution | Marie-Agnès Coutellec | Lauris Evariste, Fabrice Martin-Laurent | 2023-12-19 10:32:45 | View |
21 May 2024
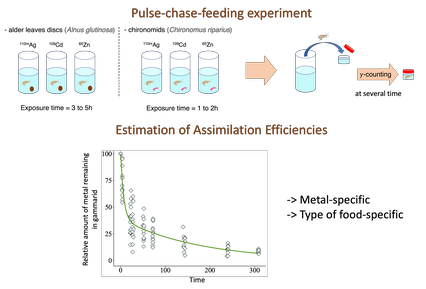
Assimilation efficiencies and elimination rates of silver, cadmium and zinc accumulated by trophic pathway in Gammarus fossarumOphélia Gestin, Christelle Lopes, Nicolas Delorme, Laura Garnero, Olivier Geffard and Thomas Lacoue-Labarthe https://doi.org/10.1101/2023.07.14.549054Food type influences dietary metal uptake and elimination in Gammarus fossarumRecommended by Patrice Couture based on reviews by Davide Anselmo Luigi Vignati and Valentin Geslin based on reviews by Davide Anselmo Luigi Vignati and Valentin Geslin
Given their narrow associations with human civilization, including urban, agricultural and industrial settings, freshwater systems worldwide are primary recipients of contaminants from anthropogenic origins, threatening biodiversity (Dudgeon 2019). Freshwater invertebrates are typically abundant in these environments. They are easily sampled, and several species can also be raised in the laboratory. Furthermore, they have the propensity to accumulate contaminants from their environments through both aqueous and dietary routes. These traits make them ideally suited as bioindicators of environmental contamination and for the study of the mechanisms of contaminant uptake and effects. Therefore, over the last decades, several studies have investigated the bioaccumulation and toxicity of a wide range of organic and inorganic contaminants. Knowledge of the relative importance of the aqueous and dietary exposure routes is key to understanding the processes involved in contaminant uptake and organismal and ecological consequences. Although the mechanisms of aqueous uptake have received much attention in recent literature, those associated with dietary uptake are far less known. This is the case for species commonly used for biomonitoring environmental contamination such as the amphipod Gammarus fossarum, and for metals of major concern for the Water Framework Directive (WFD) such as Ag, Cd and Zn. To address these knowledge gaps, Gestin et al (2024) investigated the assimilation efficiency (AE) of Ag, Cd and Zn from two contrasting types of food, one plant (alder leaves) and one invertebrate (Chironomus riparius larvae) for gammarids using a pulse-chase-feeding method in a laboratory setting. Food was radiolabeled and fed for a short period to gammarids (3 to 5 hours for alder leaves and 1 hour for chironomid larvae), after which they were left to depurate for 14 days, during which period they were fed with uncontaminated alder leaves. During the depuration period, gammarids were monitored to follow radioactivity using a gamma counter. A nonlinear least squares modelling approach was used to estimate assimilation efficiencies and elimination rates of the metals from each food source. References Dudgeon, D. (2019). Multiple threats imperil freshwater biodiversity in the Anthropocene. Current Biology 29(19):R960-R967. https://doi.org/10.1016/j.cub.2019.08.002 Gestin, O., Lopes, C., Delorme, N., Garnero, L., Geffard, O., Lacoue-Labarthe, T. (2024). Assimilation efficiencies and elimination rates of silver, cadmium and zinc accumulated by trophic pathway in Gammarus fossarum. bioRxiv, 2023.07.14.549054, ver.4 peer-reviewed and recommended by Peer Community In Ecotoxicology and Environmental Chemistry. https://doi.org/10.1101/2023.07.14.549054 | Assimilation efficiencies and elimination rates of silver, cadmium and zinc accumulated by trophic pathway in *Gammarus fossarum* | Ophélia Gestin, Christelle Lopes, Nicolas Delorme, Laura Garnero, Olivier Geffard and Thomas Lacoue-Labarthe | <p>To improve the assessment of metal toxicity in aquatic organisms, it is important to consider the different uptake pathways (i.e. trophic or aqueous). The bioaccumulation of dissolved metals such as Cd and Zn in gammarids is beginning to be wel... |  | Aquatic ecotoxicology, Bioaccumulation/biomagnification | Patrice Couture | 2023-07-15 10:27:34 | View | |
03 Jul 2024
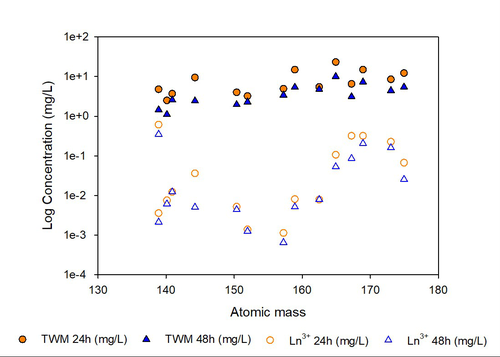
Ecotoxicity of lanthanides to Daphnia magna: insights from elemental behavior and speciation in a standardized test mediumDavide A.L. Vignati, Loïc Martin, Laurence Poirier, Aurore Zalouk-Vergnoux, Chantal Fouque, Clément Bojic, Christophe Hissler, Carole Cossu-Leguille https://hal.univ-lorraine.fr/hal-04302491v3Lanthanide atomic mass and chemical behaviour in solution influence their solubility and ecotoxicity for Daphnia magna: Implications for risk assessment of aquatic organismsRecommended by Patrice Couture based on reviews by Carrie J. Rickwood and 1 anonymous reviewer based on reviews by Carrie J. Rickwood and 1 anonymous reviewer
The demand for lanthanides (LN) has seen a steady increase and is anticipated to continue to grow. Due to their unique properties, they have become essential in key components of new technologies, such as batteries, wind turbines, electronic components and other devices needed to facilitate energy transition away from fossil fuels. These elements are also increasingly used in a range of new technologies, including medical applications and telecommunication. In this context, the concentrations of lanthanides are expected to increase in freshwater environments (Gwenzi et al., 2018). Our limited knowledge about the risk that they pose to organisms limits our ability to develop guidelines for environmental protection. Research on this issue has so far been hindered by the peculiar properties of lanthanides, that tend to form insoluble precipitates when added in standard ecotoxicological test media (Blinova et al., 2018). This and other challenges of studying lanthanide toxicity were addressed in this in-depth study that leaves few stones unturned. The study by Vignati and colleagues (2024) is the first to investigate the acute toxicity of all LN, with the exception of promethium, a radioactive element, on Daphnia magna, a model test species, following the ISO 6341 (2012) norm. The authors designed their study to generate data useable for the development of risk assessment guidelines for the LN series and to generate data-based recommendations for future studies on LN ecotoxicity. They exposed daphnids to nine to ten dilutions of all tested LN in a medium and carried out 48-hour acute immobilization assays. Initial and final pH was measured along with concentrations of LN in the test solutions sampled at various intervals by ICP-MS. This data allowed calculation of LN speciation, performed using VisualMinteq software. Effect concentrations were also calculated using different metrics based on initial (nominal), time-averaged or modelled LN3+ exposure concentrations. In their multi-faceted investigation, the authors reported several observations that clearly contribute to a better understanding of the ecotoxicity of LN to aquatic organisms and provide useful advice for future studies, briefly summarized here. Proper characterization of exposure concentrations is a key in any ecotoxicological study. Their project shows that even for a short, 48 h exposure, LN concentrations decrease due to a combination of precipitation and, possibly, adsorption. The concentration decrease was inversely proportional to the LN atomic mass, which may reduce the analytical requirements for future studies using the same test medium. The addition of LN to the test medium also modified pH and a detailed hypothesis is formulated to explain this phenomenon that has implications for ecotoxicological endpoints. Conclusions on LN ecotoxicity drawn in this study are based on experimental data and on extensive thermodynamic speciation modeling. The values of EC50 presented in the study varied by several order of magnitude depending on the chosen exposure metric, underscoring the urgent need for consensus-building on this issue across the research community. The authors also provide a comparison of their conclusions on EC50 values for daphnids with the limited data available in the literature, further validating their data with cautions carefully laid out about experimental design. The paper concludes with a list of seven caveats that should be considered both for regulators who will want to use the data presented in the paper for environmental LN concentrations regulations and for future studies. These caveats highlight the importance of considering LN speciation and chemical behavior during ecotoxicity assays, their influence on exposure concentrations, and their importance for risk assessment. They also reiterate that since LN concentrations in filtered water collected in the field are not directly comparable to EC50 values derived from laboratory studies using total or free LN3+ concentrations, an effort must be made to harmonize the methods of LN concentration measurements in field and laboratory studies. Overall, this paper may be one of the most rigorous studies in the current literature about LN ecotoxicity in freshwater systems. In its approach, it sets a precedent for future studies aiming at generating EC50 values or other toxicological endpoints of inorganic contaminants. The paper, carefully reviewed by Carrie Rickwood and by an anonymous reviewer, is a major contribution towards our understanding of LN ecotoxicity. Gwenzi, W., Mangori, L., Danha, C., Chaukura, N, Dunjana, N., Sanganyado, E. (2018). Sources, behaviour, and environmental and human health risks of high technology rare earth elements as emerging contaminants. Sci. Total Environ., 636:299-313. https://doi.org/10.1016/j.scitotenv.2018.04.235 ISO. (2012). Water quality — Determination of the inhibition of the mobility of Daphnia magna Straus (Cladocera, Crustacea) — Acute toxicity test (norm 6341). https://www.iso.org/standard/54614.html Vignati, D.A.L., Martin, L.A., Poirier, L., Zalouk-Vergnoux, A., Fouque, C., Clément, B., Hissler, C., Cossu-Leguille, C. (2024). Ecotoxicity of lanthanides to Daphnia magna: insights from elemental behavior and speciation in a standardized test medium. Ver.3 peer-reviewed and recommended by Peer Community In Ecotoxicology and Environmental Chemistry. https://hal.science/LIEC-UL/hal-04302491v3 | Ecotoxicity of lanthanides to *Daphnia magna*: insights from elemental behavior and speciation in a standardized test medium | Davide A.L. Vignati, Loïc Martin, Laurence Poirier, Aurore Zalouk-Vergnoux, Chantal Fouque, Clément Bojic, Christophe Hissler, Carole Cossu-Leguille | <p>Lanthanides (LNs) are a group of 15 elements with steadily increasing economical importance due to their multiple uses in technologies essential for sustainable ecological, digital and energetic transitions. Although knowledge on LN ecotoxicolo... |  | Aquatic ecotoxicology, Chemical speciation | Patrice Couture | 2023-11-23 15:16:50 | View | |
09 Dec 2022

Soot and charcoal as reservoirs of extracellular DNAStanislav Jelavic, Lisbeth Garbrecht Thygesen, Valerie Magnin, Nathaniel Findling, Sascha Müller, Viktoriia Meklesh, Karina Krarup Sand https://doi.org/10.26434/chemrxiv-2021-9pz8c-v5New insights into eDNA sorption onto environmental carbonaceous materialsRecommended by Pierre Labadie based on reviews by Jérôme Duval and 1 anonymous reviewerIn recent years, the use of environmental DNA (eDNA) to investigate biodiversity has gained considerable interest (Thomsen and Willerslev, 2015; Mauvisseau et al., 2022). It allows for the indirect detection of species but it requires a sound understanding of eDNA behaviour and persistence in the environment. This is, however, a complex task because eDNA may be found in several states (e.g., dissolved, adsorbed, intracellular or intraorganellar), which display specific decay rates controlled by environmental factors (Harrisson et al., 2019; Mauvisseau et al. 2022). In the environment, dissolved DNA may interact with the surfaces of various sorbents, including mineral and organic particles/colloids. Current knowledge on eDNA sorption suggests that eDNA–sorbent interactions are controlled by electrostatics as well as inner-sphere complex formation (Mauvisseau et al., 2022). In this context, the work undertaken by Jelavic et al. (2022), focused on the adsorption of eDNA by lesser-investigated carbonaceous materials (CMs), namely soot and charcoal, as common non-mineral environmental surfaces. The authors aimed to study the adsorption capacity of soot and charcoal surfaces with respect to eDNA, in relation to solution parameters (i.e., pH, ionic strength, concentration/type of cations), time and eDNA length, under both non‐equilibrium and equilibrium conditions. Using such an approach, Jelavic et al. demonstrated the large adsorption capacities of CMs and the strong binding of DNA to these sorbents. The authors did not provide definitive conclusions on the mechanisms of eDNA sorption onto CMs. However, they provided new elements suggesting that, along with electrostatic interactions, hydrophobic interactions might play an important role in the adsorption of eDNA to CMs such as soot and charcoal. Altogether, the results presented in this paper highlight the relevance of CMs as sources of biodiversity information. In addition, it is likely that those results will also prove useful for the community to improve protocols for eDNA extraction from environmental samples that contain high fractions of CMs, e.g. urban soils. References Harrison JB, Sunday JM, Rogers SM (2019) Predicting the fate of eDNA in the environment and implications for studying biodiversity. Proceedings of the Royal Society B: Biological Sciences, 286, 20191409. https://doi.org/10.1098/rspb.2019.1409 Jelavic S, Thygesen LG, Magnin V, Findling N, Müller S, Meklesh V, Sand KK (2022) Soot and charcoal as reservoirs of extracellular DNA. ChemRxiv, ver. 5 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://doi.org/10.26434/chemrxiv-2021-9pz8c-v5 Mauvisseau Q, Harper LR, Sander M, Hanner RH, Kleyer H, Deiner K (2022) The Multiple States of Environmental DNA and What Is Known about Their Persistence in Aquatic Environments. Environmental Science & Technology, 56, 5322–5333. https://doi.org/10.1021/acs.est.1c07638 Thomsen PF, Willerslev E (2015) Environmental DNA – An emerging tool in conservation for monitoring past and present biodiversity. Biological Conservation, 183, 4–18. https://doi.org/10.1016/j.biocon.2014.11.019 | Soot and charcoal as reservoirs of extracellular DNA | Stanislav Jelavic, Lisbeth Garbrecht Thygesen, Valerie Magnin, Nathaniel Findling, Sascha Müller, Viktoriia Meklesh, Karina Krarup Sand | <p style="text-align: justify;">The vast potential of using sediment adsorbed DNA as a window to past and present biodiversity rely on the ability of solid surfaces to adsorb environmental DNA. However, a comprehensive insight into DNA adsorption ... |  | Analytical Chemistry, Environmental chemistry, Environmental monitoring | Pierre Labadie | Anonymous, Jérôme Duval | 2022-04-13 16:08:36 | View |
24 Mar 2023
Identifying pesticide mixtures at country-wide scaleMilena Cairo, Anne-Christine Monnet, Stéphane Robin, Emmanuelle Porcher, Colin Fontaine https://hal.science/hal-03815557An original approach for the identification of relevant pesticides mixtures at nationwide scaleRecommended by Pierre Labadie based on reviews by Patrice Couture and Clémentine FRITSCHOver the last decades, pesticides have been massively used in agriculture and their impacts on both the environment and human health are a major growing concern (Humann-Guilleminot et al., 2019; 2019 Boedeker et al., 2020). Improving the prediction of wildlife exposure to pesticides and the associated impacts on ecosystems is therefore crucial. In general, ecotoxicological studies addressing the effects of pesticides include compounds that are selected based on general use over large areas (e.g. regions, country) or specific crop types. Such a selection does not necessarily reflect the mixtures to which species of wildlife are exposed in a particular ecosystem. In this context, Cairo et al. (2023) present an original approach to identify relevant mixtures of current-use pesticides. Their approach relies on public data concerning pesticide sales and cropping, available at a nationwide scale in France and at a relatively high resolution (i.e. postcode of the buyer). Based on a number of clearly exposed and discussed assumptions (e.g. “pesticides were used in the year of purchase and in the postcode of purchase”), their approach allowed for identifying 18 groups that were discriminated by a reduced number of pesticides. Some compounds were found in most or all groups and were termed “core substances” (e.g. deltamethrin and lambda-cyhalothrin). Other compounds, however, were associated with a limited number of groups and termed “discriminant substances” (e.g. boscalid and epoxiconazole). The authors identified groups of molecules that are probably associated with the same mixtures, which warrants the investigation of potential synergetic effects. In addition, their approach allowed for the identification of areas where aquatic biota may be exposed to similar mixtures, which is might prove of interest to further investigate in situ the actual impacts of pesticide mixtures on ecosystems. Note that the approach taken by the authors might be applied by others in other countries, provided a database of pesticide sales is available. REFERENCES Boedeker W, Watts M, Clausing P, Marquez E (2020) The global distribution of acute unintentional pesticide poisoning: estimations based on a systematic review. BMC Public Health, 20, 1875. https://doi.org/10.1186/s12889-020-09939-0 Cairo M, Monnet A-C, Robin S, Porcher E, Fontaine C (2023) Identifying pesticide mixtures at country-wide scale. HAL, ver. 2 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://hal.science/hal-03815557 Humann-Guilleminot S, Tassin de Montaigu C, Sire J, Grünig S, Gning O, Glauser G, Vallat A, Helfenstein F (2019) A sublethal dose of the neonicotinoid insecticide acetamiprid reduces sperm density in a songbird. Environmental Research, 177, 108589. https://doi.org/10.1016/j.envres.2019.108589 | Identifying pesticide mixtures at country-wide scale | Milena Cairo, Anne-Christine Monnet, Stéphane Robin, Emmanuelle Porcher, Colin Fontaine | <p style="text-align: justify;">Wild organisms are likely exposed to complex mixtures of pesticides owing to the large diversity of substances on the market and the broad range agricultural practices. The consequences of such exposure are still po... | Environmental pollution, Environmental risk assessment, Method standardization, Other | Pierre Labadie | Clémentine FRITSCH, Patrice Couture | 2022-10-14 17:13:06 | View | |
18 Jan 2022
Machine learning models based on molecular descriptors to predict human and environmental toxicological factors in continental freshwaterRémi Servien, Eric Latrille, Dominique Patureau, Arnaud Hélias https://doi.org/10.1101/2021.07.20.453034Predicting characterization factors of chemical substances from a set of molecular descriptors based on machine learning algorithmsRecommended by Sandrine CHARLES based on reviews by Patrice Couture, Sylvain Bart, Dominique Lamonica and 2 anonymous reviewers based on reviews by Patrice Couture, Sylvain Bart, Dominique Lamonica and 2 anonymous reviewers
Today, thousands of chemical substances are released into the environment because of human activities. It is thus crucial to identify all relevant chemicals that contribute to toxic effects on living organisms, also potentially disturbing the community functioning and the ecosystem services that flow from them. Once identified, chemical substances need to be associated with ecotoxicity factors. Nevertheless, getting such factors usually requires time-, resources- and animal-costly experiments that it should be possible to avoid. In this perspective, modelling approaches may be particularly helpful if they rely on easy-to-obtain information to be used as predictive variables. Within this context, the paper of Servien et al. (2022) illustrates the use of machine learning algorithms to predict toxicity and ecotoxicity factors that were missing for a collection of compounds. Their modelling approach involve a collection of molecular descriptors as input variables. A total of 40 molecular descriptors were extracted from the TyPol database (Servien et al., 2014) as those describing the best how organic compounds behave within the environment. These molecular descriptors also have the advantage to be easily quantifiable for new chemical substances under evaluation. The performances of the proposed models were systematically checked and compared to the classical linear partial least square method, based on the calculation of the absolute error (namely, the difference between prediction and true value). This finally led to different best models (that is associated to the lowest median absolute error) according to the classification of the 526 compounds comprised in the TyPol database in five clusters. These five clusters of different sizes gather chemical substances with different but specific molecular characteristics, also corresponding to different estimates of the characterization factors both in their median and within-variability. In a final step, predictions of characterization factors were performed for 102 missing values in the USEtox® database (Rosenbaum et al., 2008) but also referenced in TyPol. This paper highlights that the molecular descriptors that explain the most the toxicity of the chemical substances in each cluster strongly differ. Nevertheless, these predictions, whatever the cluster, appear precise enough to be considered as relevant despite everything. As a conclusion, this paper is a promising proof-of-concept in using machine learning modelling to go beyond some constraints around the toxicity evaluation of chemical substances, especially handling non-linearities and data-demanding calculations, in in an ever-changing world that is gradually depleting its resources without sufficient concern for the short-term risks to the environment and human health. References Rosenbaum RK, Bachmann TM, Gold LS, Huijbregts MAJ, Jolliet O, Juraske R, Koehler A, Larsen HF, MacLeod M, Margni M, McKone TE, Payet J, Schuhmacher M, van de Meent D, Hauschild MZ (2008) USEtox—the UNEP-SETAC toxicity model: recommended characterisation factors for human toxicity and freshwater ecotoxicity in life cycle impact assessment. The International Journal of Life Cycle Assessment, 13, 532. https://doi.org/10.1007/s11367-008-0038-4 Servien R, Latrille E, Patureau D, Hélias A (2022) Machine learning models based on molecular descriptors to predict human and environmental toxicological factors in continental freshwater. bioRxiv, 2021.07.20.453034, ver. 6 peer-reviewed and recommended by Peer Community in Ecotoxicology and Environmental Chemistry. https://doi.org/10.1101/2021.07.20.453034 Servien R, Mamy L, Li Z, Rossard V, Latrille E, Bessac F, Patureau D, Benoit P (2014) TyPol – A new methodology for organic compounds clustering based on their molecular characteristics and environmental behavior. Chemosphere, 111, 613–622. https://doi.org/10.1016/j.chemosphere.2014.05.020 | Machine learning models based on molecular descriptors to predict human and environmental toxicological factors in continental freshwater | Rémi Servien, Eric Latrille, Dominique Patureau, Arnaud Hélias | <p style="text-align: justify;">It is a real challenge for life cycle assessment practitioners to identify all relevant substances contributing to the ecotoxicity. Once this identification has been made, the lack of corresponding ecotoxicity facto... | Aquatic ecotoxicology, Ecosystem Health, Environmental pollution, Modelling | Sandrine CHARLES | 2021-07-21 09:09:50 | View |